Heme Biosynthesis and Heme Oxygenase Pathways
The Heme biosynthetic pathway represents one of the most essential metabolic pathways in living organisms, providing the precursor for hemoglobin. Heme is synthesized by eight enzymatic steps, starting with condensation of glycine and succinyl CoA to form d-aminolevulinic acid (ALA) by d -aminolevulinic acid synthase (ALAS)1. Two molecules of ALA are then condensed to porphobilinogen (PBG) by ALA dehydratase (ALAD)2. Four molecules of PBG are subsequently used to form the unstable hydroxymethylbilane (HMB) by porphobilinogen deaminase (PBGD)3 followed by the cyclization to uroporphyrinogen III (Uro III) by uroporphyrinogen III synthase (UROS)4. Hydroxymethylbilane will also spontaneously cyclize to uroporphyrinogen I5. The synthesis of heme continues by decarboxylation of the four acetic side chains to methyl groups by uroporphyrinogen III decarboxylase (UROD)6 to form coproporphyrinogen III (Copro III). The synthesis is continued by coproporphyrinogen III oxidase (CPO)7, 8 with the formation of protoporphyrinogen IX. Next, protoporphyrinogen oxidase (PPO)9, 10 mediates a six-electron oxidation forming protoporphyrin IX (PPIX). The final product, heme, is formed by ferrochelatase (FC)11, which mediates the insertion of ferrous iron.
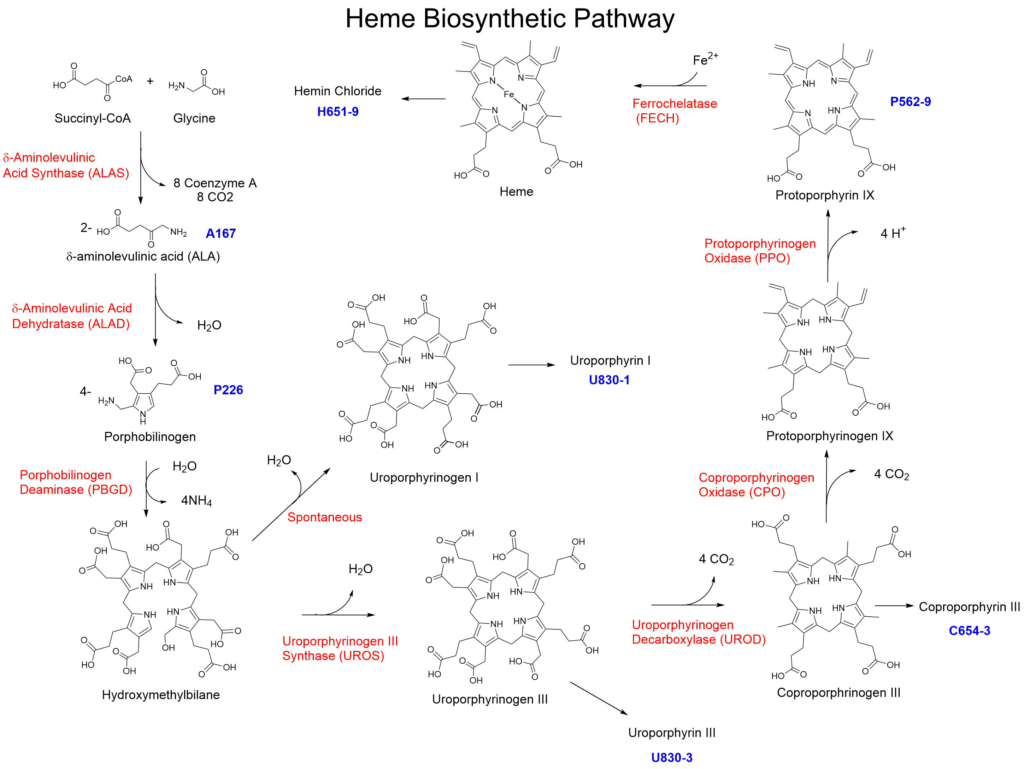
Frontier Specialty Chemicals offers several of the compounds along this pathway to assist in research into this pathway. These materials are also used to identify which enzymatic deficiencies are responsible for elevated levels of the intermediates in individuals suspected to have Porphyria disease. It should be noted that the porphyrinogen intermediates are unstable to isolation and will begin to spontaneously oxidize in air to the corresponding porphyrin analogs. To provide the highest quality of material, Frontier Specialty Chemicals provides the porphyrins corresponding to the porphyrinogens in the pathway (Uropophyrin III, Coproporphyrin III and Protoporphyrin IX).
Heme Degradation and Bilirubin Elimination Pathway
The heme oxygenase (HO)12 is the rate limiting enzyme responsible for the degradation of heme. The enzyme converts heme into carbon monoxide (CO) iron and biliverdin. Biliverdin is then converted to bilirubin via biliverdin reductase (BR)13. Bilirubin is then transported to the liver and is converted to bilirubin diglucuronide via UDP glucuronotransferase (UGT1A1)14 and is eliminated as a component of bile. Blue light is also capable of isomerizing bilirubin present in the skin to a form capable of direct elimination.
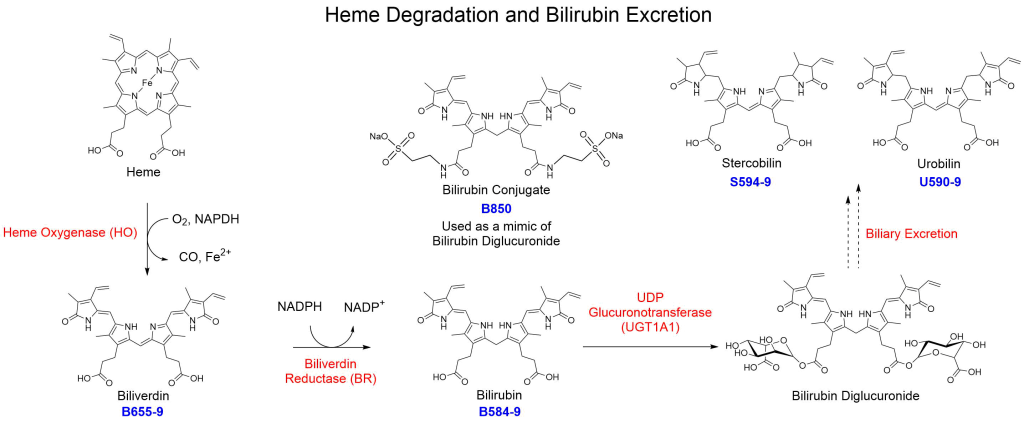
| Catalog Number: B655-9 Biliverdin | Catalog Number: B584-9 Bilirubin | Catalog Number: U590-9 Urobilin |
| Catalog Number: S594-9 Stercobilin | Catalog Number: B850 Bilirubin Conjugate |
Frontier does not sell the diglucuronide but has developed bilirubin conjugate (B850). The ditaurate form behaves like the diglucuronide within analytical testing methods. Bilirubin diglucuronide is converted via various methods within the gut microbiome and is ultimately eliminated in the stool as bilirubin, stercobilin or as urobilin.
References
1. Ferreira, G.C., Gong, J. 5-Aminolevulinate synthase and the first step of heme biosynthesis. J Bioenerg Biomembr 27, 151–159 (1995). https://doi.org/10.1007/BF02110030
2. Jaffe, E.K. Porphobilinogen synthase, the first source of Heme’s asymmetry. J Bioenerg Biomembr 27, 169–179 (1995). https://doi.org/10.1007/BF02110032
3. Grandchamp, B., De Verneuil, H., Beaumont, C., Chretien, S., Walter, O. and Nordmann, Y. (1987), Tissue‐specific expression of porphobilinogen deaminase. European Journal of Biochemistry, 162: 105-110. doi:10.1111/j.1432-1033.1987.tb10548.x
4. Battersby, A., Fookes, C., Matcham, G. et al. Biosynthesis of the pigments of life: formation of the macrocycle. Nature 285, 17–21 (1980). https://doi.org/10.1038/285017a0
5. Paul R. Ortiz de Montellano (2008). “Hemes in Biology”. Wiley Encyclopedia of Chemical Biology. John Wiley & Sons. doi:10.1002/9780470048672.wecb221
6. Straka, J.G., Kushner, J.P Purification and characterization of bovine hepatic uroporphyrinogen decarboxylase. Biochemistry 1983, 22, 20, 4664-4672. https://doi.org/10.1021/bi00289a009
7. T. Yoshinaga, S. Sano Coproporphyrinogen oxidase: I. Purification, properties, and activation by phospholipids. J. Biol. Chem., 255 (10) (1980), pp. 4722-4726.
8 T. Yoshinaga, S. Sano Coproporphyrinogen oxidase: II. Reaction mechanism and role of tyrosine residues on the activity. J. Biol. Chem., 255 (10) (1980), pp. 4727-4731.
9. Dailey HA. 1990. Conversion of coproporphyrinogen to protoheme in higher eukaryotes and bacteria: Terminal three enzymes. In Biosynthesis of heme and chlorophylls (ed. Dailey HA), pp. 123–161. McGraw-Hill, New York.
10. Dailey, T. A. & Dailey, H. A. Human protoporphyrinogen oxidase: Expression, purification, and characterization of the cloned enzyme. Protein Sci. 5, 98–105 (1996). https://doi.org/10.1002/pro.5560050112
11. Lecerof, D.; Fodje, M.; Hansson, A.; Hansson, M.; Al-Karadaghi, S. (March 2000). “Structural and mechanistic basis of porphyrin metallation by ferrochelatase”. Journal of Molecular Biology. 297 (1): 221–232. doi:10.1006/jmbi.2000.3569
12. Kikuchi G, Yoshida T, Noguchi M (2005). “Heme oxygenase and heme degradation”. Biochem. Biophys. Res. Commun. 338 (1): 558–567. doi:10.1016/j.bbrc.2005.08.020
13. Ahmad Z, Salim M, Maines MD (Mar 2002). “Human biliverdin reductase is a leucine zipper-like DNA-binding protein and functions in transcriptional activation of heme oxygenase-1 by oxidative stress”. The Journal of Biological Chemistry. 277 (11): 9226–32. doi:10.1074/jbc.M10823920
14. Jansen, P.L.M., Mulder, G.J., Burchell, B. and Bock, K.W. (1992), New developments in glucuronidation research: Report of a workshop on “Glucuronidation, its role in health and disease”. Hepatology, 15: 532-544. doi:10.1002/hep.1840150328
15. McDonagh, A.F., Palma, L.A., Lightner, D.A., “Blue Light and bilirubin excretion” Science, 1980, 208, 145-151. DOI: 10.1126/science.7361112